Cross-Modal Knowledge Distillation from Spatial Transcriptomics to Histology

Abstract
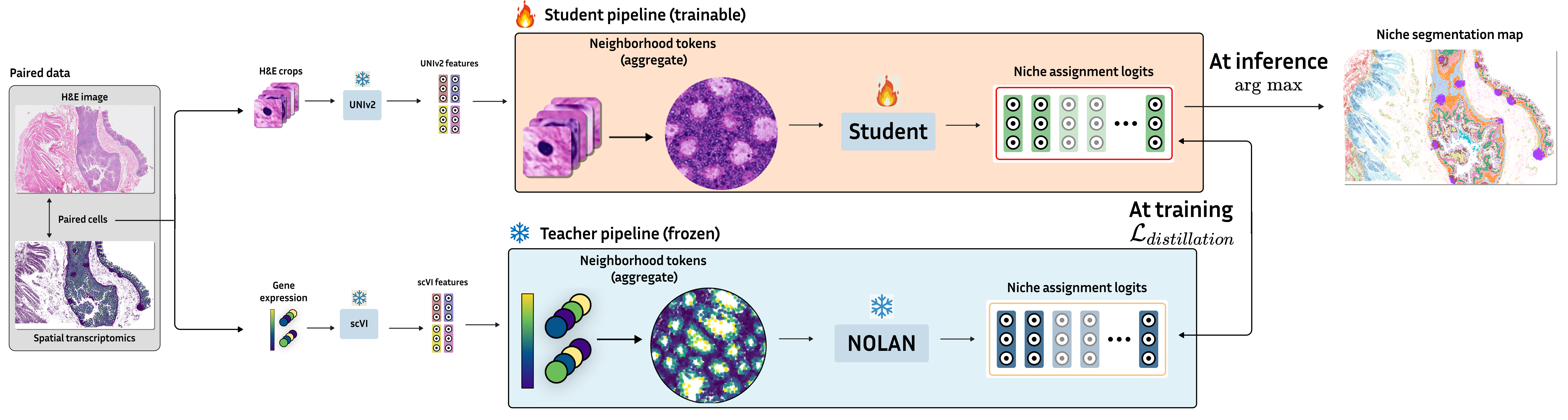
Spatial transcriptomics provides a molecularly rich description of tissue organization, enabling unsupervised discovery of tissue niches --- spatially coherent regions of distinct cell-type composition and function that are relevant to both biological research and clinical interpretation. However, spatial transcriptomics remains costly and scarce, while H&E histology is abundant but carries a less granular signal. We propose to leverage paired spatial transcriptomics and H&E data to transfer transcriptomics-derived niche structure to a histology-only model via cross-modal distillation. Across multiple tissue types and disease contexts, the distilled model achieves substantially higher agreement with transcriptomics-derived niche structure than unsupervised morphology-based baselines trained on identical image features, and recovers biologically meaningful neighborhood composition as confirmed by cell-type analysis. The resulting framework leverages paired spatial transcriptomic and H&E data during training, and can then be applied to held-out tissue regions using histology alone, without any transcriptomic input at inference time.
BibTeX
@misc{hizmi2026cross,
title={Cross-Modal Knowledge Distillation from Spatial Transcriptomics to Histology},
author={Arbel Hizmi and Artemii Bakulin and Shai Bagon and Nir Yosef},
year={2026},
eprint={2604.09076},
archivePrefix={arXiv},
primaryClass={cs.CV},
url={https://arxiv.org/abs/2604.09076}
}